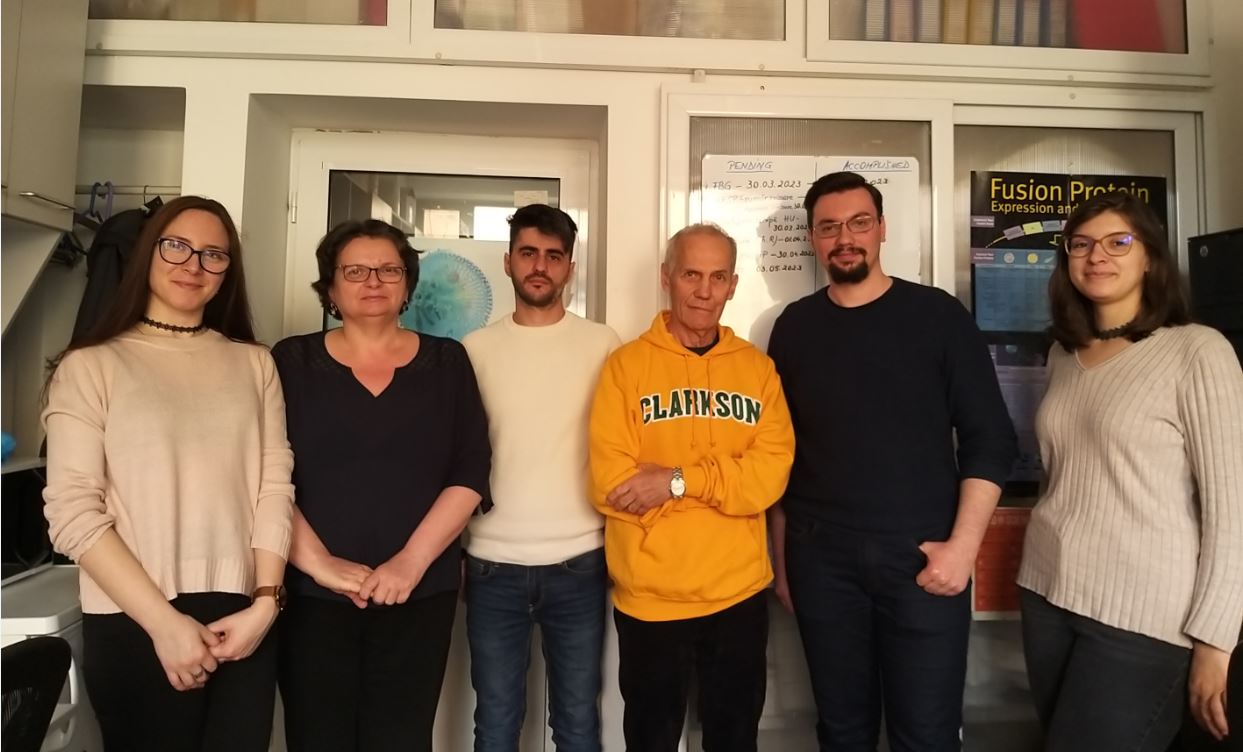

Our central research topic is the study of structure-function relationships in signaling enzymes, with a focus on protein tyrosine phosphatases. The aim is to contribute to the understanding of how their structural features are correlated with specific signaling functions. To this end, signaling enzymes are studied from several directions:
The combination of results thus obtained in this way is further used to shed light on the signaling mechanism and overall functional role of the given enzyme.
We have good experience and we are currently involved in the production, isolation and purification of recombinant proteins, expressed in both prokaryotic and eukaryotic systems. Our research activity is carried out through tools of molecular biology (recombinant DNA, site-directed mutagenesis, (RT)-PCR, Western blot, immunoprecipitation, etc.), spectroscopic analysis (UV-VIS and fluorescent spectrophotometry), cell biology, protein crystallization and enzyme kinetics.
Our ongoing research projects are:
Current research directions:
(a) expression, purification and study of the mechanism of action of some proteins involved in synaptic transmission;
(b) creation of new conjugates based on afibodies with a potential therapeutic role in different forms of leukemia;
(c) biochemical study of tau protein acetylation with applicability in neurodegenerative diseases;
(d) study of signaling mechanisms involved in various cancer forms
External website: full link: https://www.nipne.ro/proiecte/pn3/20-projects.html
External website: full link: https://www.nipne.ro/proiecte/pn3/9-projects.html
External website: full link: https://www.icbp.ro/static/en/en-networking_grants-grants national_grants/pcca1contract_792012.html
External website: full link: https://www.nipne.ro/proiecte/pn2/149-projects.html
Outstanding researchers who have made a significant contribution to the advancement of knowledge in their field, demonstrating the importance of the solid foundation provided by the Department of Enzymology in their professional training.
1995-2005

During this period, the Department of Enzymology at theInstitute of Biochemistry was characterised by innovative research that had a profound impact on the scientific field. The research team made remarkable progress in various fields of biology and medicine, contributing significantly to the understanding of the molecular mechanisms of diseases, the development of innovative genetic therapies and the elucidation of the structure of proteins involved in pathological processes. One of the most notable achievements of this period was the first successful cloning in Romania, made possible by the diligent work and close collaboration of Radu Aricescu, Valeriu Bogdan Cișmașiu, Tudor Fulga and Ștefan Szedlacsek.
This major achievement involved the synthesis of the nucleotide sequence and its cloning into a prokaryotic expression vector, followed by the expression, purification and characterisation of the resulting protein. This collaborative effort opened new perspectives in the understanding of molecular mechanisms and advanced knowledge in structural biology. The successful cloning of a tyrosine phosphatase receptor protein was a milestone in the department's research.
These researchers demonstrated that through collaboration and perseverance, major innovations are possible and their impact is felt not only in Romania, but throughout the international scientific community. Their work opened up new avenues of research and had a global impact, collaborating with prestigious institutions around the world. Their results have been instrumental in advancing knowledge in enzymology and molecular biology, laying the foundations for future research and positively influencing the international scientific community.
2005 - 2015
Another generation of talented researchers has been involved in advanced studies to elucidate the structure and function of essential proteins using X-ray diffraction and super-resolution microscopy techniques. They participated in studies investigating the substrates of EYA3 phosphatase and how its dephosphorylation influences changes in the actin cytoskeleton, with results published in the prestigious journal Scientific Reports.
They have contributed to the understanding of the enzymatic properties and substrate specificity of phosphoketolases from several bacteria, including Lactococcus lactis, Leuconostoc mesenteroides and Pseudomonas aeruginosa. These studies have provided essential biochemical and kinetic data for the application of these enzymes in biotechnological processes.
2015 -2023
under construction
under construction
RECENT PUBLICATIONS:
1. "Breaking the cancer code: a novel DNA minicircle to disable STAT3 in ovarian cancer cells SKOV3"; Adina-Gabriela Vasilescu, Andrei-Mihai Vasilescu, Livia Elena Sima, Natalia Baran, Ștefan-Eugen Szedlacsek, Frontiers in Pharmacology; 2025; 10.3389/fphar.2025.1673427
2. "A novel cytotoxic anti-B7-H3 affibody with therapeutic potential in acute myeloid leukemia" ; Andrei-Mihai Vasilescu, Adina-Gabriela Vasilescu, Livia Elena Sima, Cristian V.A. Munteanu, Natalia Baran, Stefan-Eugen Szedlacsek; Frontiers in Pharmacology; 2025; 10.3389/fphar.2025.1684226
3. "Mn(II)-based diagnostic agents: from basic research to targeted diagnostic procedures"; Balázs Váradi, Gergő Zoltán Sajtos, Károly Brezovcsik, Zoltan Szűcs, Stefan Szedlacsek, Gábor Nagy, Gyula Tircso; Scientia et Securitas; 2024; https://doi.org/10.1556/112.2024.00208
4. Editorial: „Molecular targets in oncological and hematological disease management: innovations in precision medicine"; Adrian Bogdan Tigu, Gregory Wiedman, Stefan Eugen Szedlacsek; Frontiers in Pharmacology; 2024; DOI: 10.3389/fphar.2024.1494396
5. "Synthesis and characterization of a novel [52Mn]Mn-labelled affibody based radiotracer for HER2+ targeting"; Balázs-Váradi, Károly-Brezovcsik, Zoltán Garda, Eniko-Madarasi, Horea-Szedlacsek, Rodica-Aura Badea, Andrei-Mihai Vasilescu, Adina-Gabriela Puiu, Aura Ionescu, Livia-Elena Sima, Cristian V.A. Munteanu, Simona Călăraș, Adrienn Vagner, Dezső Szikra, Ngô Minh Toàn, Tibor Nagy, Zoltán Szűcs, Stefan Eugen Szedlacsek, Inorganic Chemistry Frontiers; 2023, 10.1039/D3QI00356F
6. "Designed Peptide Inhibitors of STEP Phosphatase-GluA2 AMPA Receptor Interaction Enhance the Cognitive Performance in Rats"; Szedlacsek HS, Bajusz D, Badea RA, Pop A, Bică CC, Ravasz L, Mittli D, Mátyás D, Necula-Petrăreanu G, Munteanu CVA, Papp I, Juhász G, Hritcu L, Keserű GM, Szedlacsek SE. J Med Chem. 2022; 10.1021/acs.jmedchem.1c01303
7. "Trojan horse treatment based on PEG-coated extracellular vesicles to deliver doxorubicin to melanoma in vitro and in vivo"; Patras L, Ionescu AE, Munteanu C, Hajdu R, Kosa A, Porfire A, Licarete E, Rauca VF, Sesarman A, Luput L, Bulzu P, Chiroi P, Tranca RA, Meszaros MS, Negrea G, Barbu-Tudoran L, Potara M, Szedlacsek S, Banciu M. Cancer Biol Ther; 2022; 10.1080/15384047.2021.2003656
8. “Comparative Analysis Between Cellular Oncogenes and Viral Oncogenes"; Biointerface Research in Applied Chemistry; Puiu, AG; Grigoras, O; Preda, MI; Constantin, M; Vasilescu, AM.; 2021; Review:10.33263/BRIAC113.99399951
9. ”Regulation of TRPM8 Channel activity by Src-mediated Tyrosine Phosphorylation”; Alexandra Manolache, Tudor Selescu, G. Larisa Maier, Mihaela Mentel, Aura Elena Ionescu, Cristian Neacșu, Alexandru Babeș, Ștefan Eugen Szedlacsek. Journal of Cellular Physiology;2020; 10.1002/jcp.29397
10. ”Analysis of EYA3 phosphorylation by Src kinase identifies residues involved in cell Proliferation”; Aura E. Ionescu, Mihaela Mentel, Cristian V.A. Munteanu, Livia E. Sima, Eliza C. Martin, Georgiana Necula-Petrareanu and Ștefan E. Szedlacsek. International Journal of Molecular Sciences, 2019; DOI: 10.3390/ijms20246307
11. ”Biological and molecular modifications induced by cadmium and arsenic during breast and prostate cancer development”; Environmental Research; Zimta, AA; Schitcu, V; Gurzau, E; Stavaru, C ; Manda, G; Szedlacsek, S; Berindan-Neagoe; 2019; 10.1016/j.envres.2019.108700
12. ”Crystal structure of a xylulose 5-phosphate phosphoketolase. Insights into the substrate specificity for xylulose 5-phosphate”, Journal of Structural Biology; Scheidig, AJ ; Horvath, D; Szedlacsek, SE; 2019; 10.1016/j.jsb.2019.04.017
13. ’’Collagen regulates the ability of endothelial progenitor cells to protect hypoxic myocardium through a mechanism involving miR-377/VE-PTP axis”; Rosca AM, Mitroi DN, Cismasiu V, Badea R, Necula-Petrareanu G, Preda MB, Niculite C, Tutuianu R, Szedlacsek S, Burlacu A. J Cell Mol Med.; 2018; DOI: 10.1111/jcmm.13712
14. ’’WDR1 is a novel EYA3 substrate and its dephosphorylation induces modifications of the cellular actin cytoskeleton”; Mentel M, Ionescu AE, Puscalau-Girtu I, Helm MS, Badea RA, Rizzoli SO, Szedlacsek SE. Sci Rep.; 2018; 10.1038/s41598-018-21155-w
15. “Phosphoketolases from Lactococcus lactis, Leuconostoc mesenteroides and Pseudomonas aeruginosa: dissimilar sequences, similar substrates but distinct enzymatic characteristics“; Petrareanu G, Balasu MC, Vacaru AM, Munteanu CV, Ionescu AE, Matei I, Szedlacsek SE. Appl Microbiol Biotechnol.; 2014; 10.1007/s00253-014-5723-6
16. “Protein tyrosine phosphatase structure-function relationships in regulation and pathogenesis“; Böhmer F, Szedlacsek S, Tabernero L, Ostman A, den Hertog J.; FEBS J. 280, 2013; 10.1007/s00253-014-5723-6
17. “Preliminary X-ray crystallographic analysis of the D-xylulose 5-phosphate phosphoketolase from Lactococcus lactis”; Petrareanu, G, Balasu, MC, Zander, U, Scheidig, AJ and Szedlacsek, S.E., Acta Cryst.; 2010; 10.1107/S174430911001732X
18. “Interface Analysis of the Complex between ERK2 and PTP-SL”; Balasu MC, Spiridon LN, Miron S, Craescu CT, Scheidig AJ, Petrescu AJ, Szedlacsek SE. PLoSOne.; 2009; 10.1371/journal.pone.0005432
19. “Analysis of Molecular Determinants of PRL-3”; Pascaru M, Tanase C, Vacaru AM, Boeti P, Neagu E, Popescu I, Szedlacsek SE.; J Cell Mol Med.; 2009;10.1111/j.1582-4934.2008.00591.x
20. “Protein tyrosine phosphatases, structure-function relationships”; Tabernero L, Aricescu AR, Jones EY, Szedlacsek SE. FEBS J.; 2008; 10.1111/j.1742-4658.2008.06251.x
21. “A microarray strategy for mapping the substrate specificity of protein tyrosine phosphatase”; Köhn M, Gutierrez-Rodriguez M, Jonkheijm P, Wetzel S, Wacker R, Schroeder H, Prinz H, Niemeyer CM, Breinbauer R, Szedlacsek SE, Waldmann H. Angew Chem Int Ed Engl.; 2007; 10.1002/anie.200701601
22. “Functional, fractal nonlinear response with application to rate processes with memory, allometry, and population genetics”; Vlad MO, Morán F, Popa VT, Szedlacsek SE, Ross J.; Proc Natl Acad Sci USA., 2007; DOI: 10.1073/pnas.0700397104
23. “Identification and specificity profiling of protein prenyltransferase inhibitors using new fluorescent phosphoisoprenoids”; Dursina B, Reents R, Delon C, Wu Y, Kulharia M, Thutewohl M, Veligodsky A, Kalinin A, Evstifeev V, Ciobanu D, Szedlacsek SE, Waldmann H, Goody RS, Alexandrov K. J Am Chem Soc.; 2006; 10.1021/ja052196e
24. “Fisher's theorems for multivariable, time-and space-dependent systems, with applications in population genetics and chemical kinetics”; Vlad MO, Szedlacsek SE, Pourmand N, Cavalli- Sforza LL, Oefner P, Ross J; Proc Natl Acad Sci USA; 2005; 10.1073/pnas.0504073102
25. “The MAM (Meprin/A5-protein/PTPmu) Domain Is a Homophilic Binding Site Promoting the Lateral Dimerization of Receptor-like Protein-tyrosine Phosphatase μ”; V.B. Cismasiu, S.A. Denes, H. Reilander, H. Michel, and S.E. Szedlacsek; J. Biol. Chem.; 2004; 10.1074/jbc.M313115200
26. “Synthesis and biological applications of a new 1,2,5-oxadiazolo[3,4-c]pyridine fluorescent marker”; M.C. Balasu, I. Costea, R. Fratila, A. Popescu, C. Draghici and S.E. Szedlacsek; Rev. Roum. Chim., 2004; https://doi.org/10.1002/chin.200502149
27. “Protein Tyrosine Phosphatase Inhibitors”; M.C. Balasu and S.E. Szedlacsek; Rev. Chim.; 2002; https://www.webofscience.com/wos/woscc/full-record/WOS:000177305200002
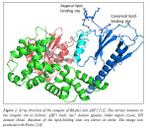
28. “Crystal structure of PTP-SL/BR7 catalytic domain, Implications for MAP kinase regulation”; S.E. Szedlacsek, A.R. Aricescu, T.A Fulga, L. Renault, A.J. Scheidig.; J. Mol. Biol.; 2001; 10.1006/jmbi.2001.4890
29. “Intramolecular interactions in protein tyrosine phosphatase RPTPμ, Kinetic evidence”; A.R Aricescu, T.A Fulga, V., Cismasiu, R.S. Goody, S.E. Szedlacsek; Biochem. Biophys. Res. Comm.; 280, 319-327; 2001;10.1006/bbrc.2000.4094
30. “Time-dependent control of metabolic systems by external effectors”; S.E. Szedlacsek, A.R. Aricescu, B.H. Havsteen; J. theor. Biol.;1996; 10.1006/jtbi.1996.0173
31. “pH-dependent hysteretic behaviour of human myeloblastin (leucocyte proteinase 3)”; A. Baici, S.E. Szedlacsek, H. Früh, B.A. Michel; Biochem. J.; 1996; 10.1042/bj3170901
32. “Esterification of oxysterols by human plasma lecithin cholesterol acyltransferase”; S.E. Szedlacsek, E. Wasowicz, H. Nishida, S.A. Hulea, F. A. Kummerow, T. Nishida, J. Biol. Chem. 1995; 10.1074/jbc.270.20.11812
33. “Kinetics of slow and tight-binding inhibitors”, S.E. Szedlacsek, R.G. Duggleby; Methods Enzymol.; 1995; 10.1016/0076-6879(95)49034-5
34. ”Steady-state analysis of the reversible closed bicyclic enzyme cascades”; VaronN, R; Havsteen, BH; Szedlacsek, SE; Garcia Moreno, M; Molina Alarcon, M; Sanchez Gracia, A; Volume: 1994; 10.1016/0020-711x(94)90108-2
35. “Kinetic analysis of reversible closed bicyclic enzyme cascades covering the whole course of the reaction”; R. Varon, B.H. Havsteen, M. Molina-Alarcon, S.E. Szedlacsek, F. Garcia-Canovas; Int. J. Biochem;1994; 10.1016/0020-711x(94)90108-2
36. “Response coefficients of interconvertible enzyme cascades towards effectors that act on one or both modifier enzymes”; S. E. Szedlacsek, M.-L. Cardenas, A. Cornish-Bowden, Eur. J. Biochem.; 1992; DOI: 10.1111/j.1432-1033.1992.tb16699.x
37. “Egg-white avidin purification by affinity elution from CM-cellulose”; C. Borza, B. Borza, F. Nitu, S. E. Szedlacsek, Rev. Roum. Biochim.;1992
38. “Very large response coefficients in interconvertible enzyme cascades”; A. Cornish-Bowden, S. E. Szedlacsek; Biomed. Biochim. Acta;1990
39. “Progress-curve equations for reversible enzyme-catalysed reactions inhibited by tight- binding inhibitors”; S.E. Szedlacsek, V. Ostafe, R.G. Duggleby, M. Serban, M.O. Vlad; Biochem. J.;265, 1990; 10.1042/bj2650647
40. “Purification of aprotinin from bovine lung extracts” (in romanian); H.D. Schell, S.E. Szedlacsek si V. Ostafe; Stud. cercet. Biochim.; 1990
The main idea of this project was that by inhibiting at least one of these two interactions of GluA2, the internalization of AMPAR will be reduced and therefore the synapse resistance will be increased, thus leading to improved cognitive functions.
The project is agreed as a bilateral collaboration between IBAR and ATOMKI, and the University of Debrecen and IFIN-HH participate in this project voluntarily.
The project is agreed as a joint collaboration among IBAR, ATOMKI and UD, the latter being a cost free participant. There are two main directions envisaged by the proposed project: - receptors mapping and therapy, using an affibody against HER2 receptors, combined with an adequate radioisotope. In this respect, the specific objectives are: a) expression and purification of affibodies; b) establishing labelling procedures; c) ex vivo and/or in vivo testing of optimal compounds.
Affibody molecules are highly promising therapeutic candidates due to their advantageous features like: small size, efficient delivery, straightforward engineering towards improved formats, site-directed conjugation of payloads, possibility of GMP production by chemical synthesis or inexpensive bacterial production leading to low product costs.
Project Funded under: Human resources and Mobility in the specific programme for research, technological development and demonstration "Structuring the European Research Area" under the Sixth Framework Programme 2002-2006. PTPNET was a Training Network for young scientists in the field of protein tyrosine phosphatase (PTP) research.
Proteins are essential players of all biological processes and they are involved in practically every function performed by a living cell.
The Alexander von Humboldt Foundation promotes academic cooperation between foreign and German researchers at the highest level. Each year, the Foundation awards more than 700 research fellowships and awards to support researchers from abroad who come to Germany to work on a research project together with German researchers at a host institute. The research topic is chosen by the applicant.
The Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) is the most important independent research funding organisation in Germany. It supports both scientific and humanistic studies and aims to promote top-level research in various disciplines at universities and research institutions. The main focus is on funding projects proposed by the scientific community, with an emphasis on knowledge-driven research.
Proteins are the most important functional units of living matter: all cellular processes are governed by specific proteins possessing highly specialized functional roles. Understanding the intimate mechanisms of cell functions necessitates a detailed knowledge of the 3D-structure of proteins.
One of the most powerful techniques in modern biomedical research is the determination of the three dimensional structure of proteins. This project proposes to establish a laboratory for 3D protein structure determination using X-ray diffraction experiments.
Protein tyrosine phosphatases (PTPs) are fundamental regulatory enzymes that dephosphorylate phosphotyrosine residues and are essential components of intracellular signalling pathways in both normal and pathological conditions. There is experimental evidence that disruption of the phosphatase activity of many PTPases is involved in the pathogenesis of several congenital or acquired diseases, including diabetes, cancer, infections, autoimmune, neuronal and cardiovascular diseases.
The project focuses on characterizing the branchio-oto-renal (BOR) syndrome, an autosomal dominant genetic disorder caused by mutations in the EYA gene. The goal is to enzymatically analyze the conserved C-terminal domain of the Eya3 protein and identify physiological substrates to better understand the molecular mechanisms involved in this syndrome.
Cancer causes 1.5 million deaths a year in Europe, accounting for a quarter of all deaths. The incidence of cancer is increasing, with 20 million new cases and 10 million deaths expected in the next 20 years. Approximately 90% of deaths are due to metastases rather than primary tumours. Genetic mutations and genomic instability allow tumour cells to invade and form metastases.
This project aims to develop a highly efficient enzyme for the conversion of xylose from cellulosic wastes into high-value biosynthetic intermediates. By optimising the catalytic efficiency of phosphoketolase and designing an enzymatic reactor, the project aims to provide a sustainable method of utilising lignocellulosic residues.